Alcohol keeps eDNA at the party longer
Have you ever collected environmental DNA (eDNA) water samples and struggled with the logistics of keeping them cool? Perhaps you’ve worked in remote areas where keeping things cool isn’t easy, or you’ve been challenged to send samples to the lab quickly.
We have some good news for you!
We need to stop eDNA from leaving the party…
Once water samples are taken and filtered to capture cellular material from animals for eDNA analysis, the DNA will continue to degrade over time. This can compromise the quality of the sample and reduce the ability to detect species.
Our standard filter preservation methods have relied upon filters stored at temperatures below 4 °C. As many of our clients have experienced, this can be challenging in field situations.
How can we get better at this?
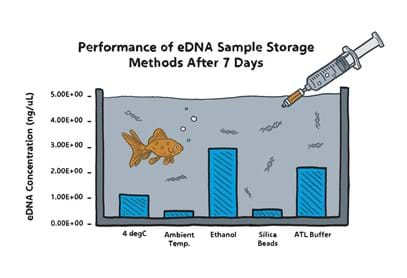
Our laboratory team have been busy assessing other methods of preservation. Using fish tank water containing goldfish, we tested various sample preservation methods, including flushing the samples with ethanol (100%); storing the filters in silica beads (desiccant) and immersing the sample in lysis buffer (used during the extraction process to break open the animal cells and release the DNA). We compared these methods to storing the samples below 4 °C as well as leaving them at room temperature.
What does this mean for our clients?
Along with the standard storage method of keeping samples cool (which continues to be an effective option for preserving eDNA), we are very excited to now be offering this ethanol sample storage method to our clients! Particularly if fieldwork is taking place in remote areas with little to no access to refrigeration or if there may be some time between collecting the samples and posting them to our lab. Provided all instructions are adhered to, we can feel very confident in our results!
If you are keen to adopt this method for your next project or have any questions please contact one of our amazing eDNA specialists, Sarah Licul, to discuss further Email: slicul@envirodna.com Ph: 9028 8753.
We aim to publish this study in the near future. Current supportive literature includes:
- Spens, J., Evans, A.R., Halfmaerten, D., Knudsen, S.W., Sengupta, M.E., Mak, S.S.T., Sigsgaard, E.E. and Hellström, M. (2017), Comparison of capture and storage methods for aqueous macrobial eDNA using an optimized extraction protocol: advantage of enclosed filter. Methods Ecol Evol, 8: 635-645. doi:10.1111/2041-210X.12683
- Hinlo R, Gleeson D, Lintermans M, Furlan E (2017) Methods to maximise recovery of environmental DNA from water samples. PLoS ONE 12(6): e0179251. https://doi.org/10.1371/ journal.pone.0179251
- Minamoto, Toshifumi; Naka, Takafumi; Moji, Kazuhiko; Maruyama, Atsushi (2015) Techniques for the practical collection of environmental DNA: filter selection, preservation, and extraction. Limnology . Jan2016, Vol. 17 Issue 1, p23-32. 10p.